-
中药红花(Carthami Flos)是菊科植物红花(Carthamus tinctorius L.)的干燥花,传统本草学著作《本草纲目》记载,红花具有活血散瘀,通经止痛的功效[1],其药材和制剂在临床上被广泛用于心脑血管疾病的预防和治疗。现代药理研究表明,其主要药效物质是以羟基红花黄色素A(hydroxysafflower yellow A,HSYA)为代表的查尔酮类化合物和以菸花苷为代表的黄酮醇类化合物,这些化合物均具有良好的心脑血管损伤保护活性[2-3]。红花药材的产量偏低,每平方千米产量仅为18.0~22.5 t[4],其中特有的HSYA[5]、红花红色素等查尔酮类成分在不同品种间差异较大[6]。由于红花中的查尔酮类成分仅特异性地存在于花冠中[7],加之体外组织培养再生率低[8]等原因,对其功能基因的研究工作一直进展缓慢。特别是对于HSYA等红花特有的有效成分,其生物合成相关的功能基因尚不完全清楚,合成通路也未被完全解析[9]。因此,用现代分子生物学技术手段以提高药效物质的含量,是提高红花品质,节约土地资源、降低制药成本的一条新途径。
短链脱氢酶/还原酶(short-chain dehydrogenases/reductases,SDR)在植物次生代谢物的生物合成中广泛参与各类碳-氧双键,碳-碳双键以及烯酮键的氧化还原催化反应。根据SDRs基因序列的特征结构,SDRs超家族可以被分为5个亚家族[10-14]。最早发现并且进行鉴定的两类主要短链还原酶命名为classical和extend,classical类的SDRs基因拥有长度约为250个氨基酸残基,被称为Extended类的SDRs基因在碳基末端因其含有多余的约100个氨基酸残基而得名。另外3种类型SDRs基因分别被命名为intermediate、complex和divergent。这些类型的SDRs基因基于其结合辅酶类型和结合催化位点的不同进行命名分类。此外,SDRs存在与传统类型不同的含有“rossmann-fold”保守结构域的氧化还原酶结构[15-18]。
黄酮类化合物起源于莽草酸途径和苯丙素生物合成途径,1个香豆酰辅酶A(coumaroyl CoA)和3个丙二酰辅酶A(malonyl CoA)在查尔酮合酶的作用下生成二氢查尔酮,然后经查尔酮异构酶催化为二氢黄酮,进一步在各类还原酶,聚合酶和糖基转移酶的作用下,生成终端次生代谢产物组合[19-21]。红花中所含的主要有效成分HSYA具有查尔酮式结构,本课题组前期研究认为:HSYA从前体物质到合成,中间存在必不可少的氧化还原过程。短链脱氢还原酶家族广泛参与植物体内次生代谢,这一类还原酶都带有相似的折叠结构以及催化位点,已有研究表明,其对苯丙烷代谢途径起重要作用[22-23],但有关红花中还原酶基因相关报道较少[24]。故笔者通过对红花转录组数据库、基因表达谱数据库以及代谢组数据库进行分析,筛选在HSYA生物合成途径的关键还原酶基因,并进行功能验证,以期揭示红花次生代谢成分生物合成途径,为定向调控红花的品质提供科学依据。
-
云南巍山红花品系(ZHH0119),采自海军军医大学药学系温室,经海军军医大学郭美丽教授鉴定为菊科植物红花(Carthamus tinctorius L.)。红花种植条件:温度恒定25 ℃,16 h光照,8 h黑暗。采集相关花与组织后迅速存放于液氮或者−80 ℃冰箱中冷冻。
-
按照Trans ZOL Plant植物总RNA提取试剂盒(北京全式金公司,中国)说明书方法提取红花花冠总RNA,按照Transtart One-Step gDNA Removal and cDNA Synthesis Super Mix逆转录试剂盒(北京全式金公司,中国)说明书方法进行cDNA第一链的合成。cDNA于−20 ℃保存。
-
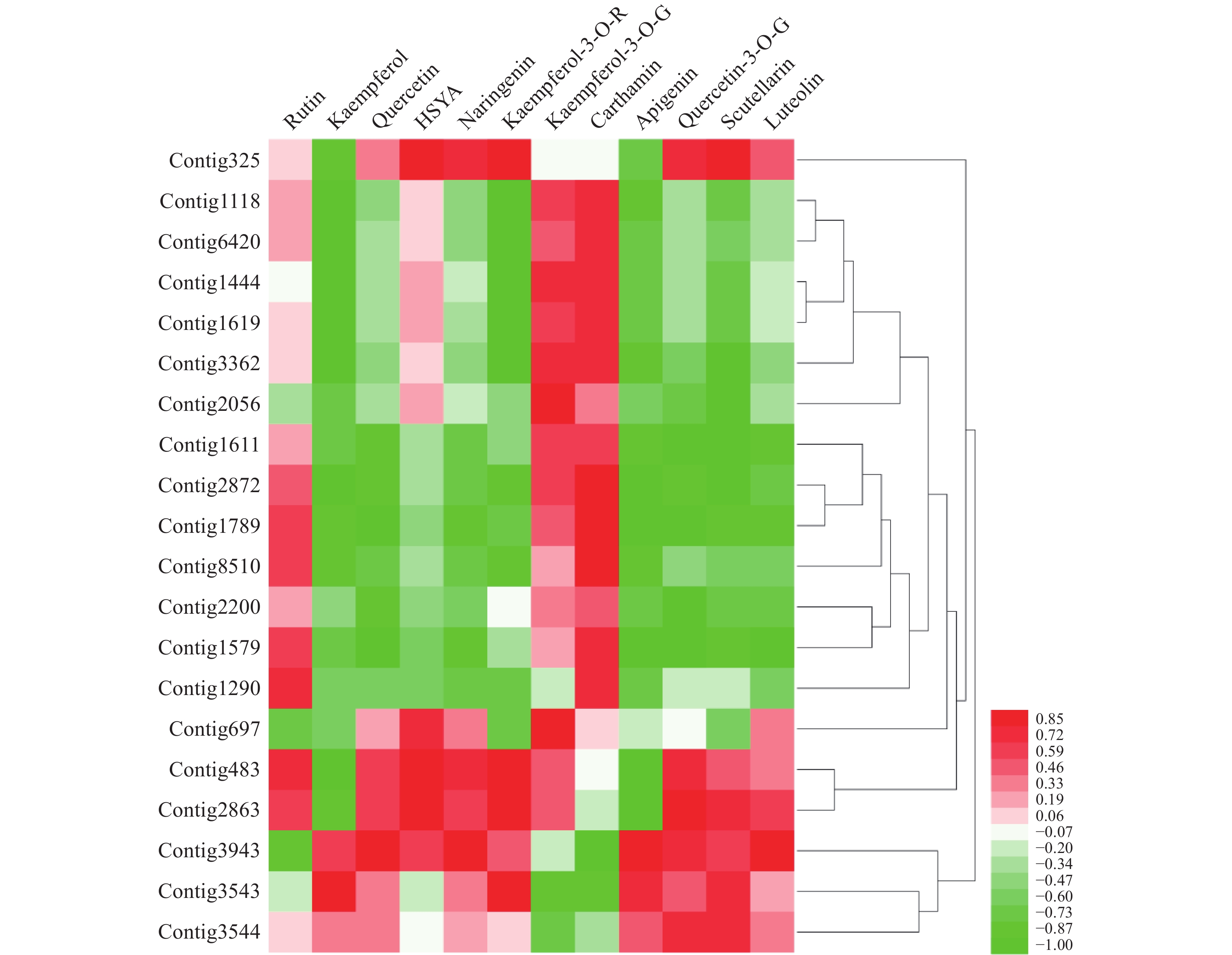
基于数据库中的基因注释以“黄酮还原酶”和“黄酮类化合物生物合成”作为关键词进行检索,筛选出其中可能与HSYA生物合成相关的还原酶基因,将筛选基因不同花期时间的表达量,将其与红花代谢组数据库中同花期的芦丁(rutin)、山柰酚(kaempferol)、槲皮素(quercetin)、HSYA、柚皮素(naringenin)、山柰酚-3-O-芸香糖苷(kaempferol-3-O-rutinoside)、山柰酚-3-O-葡萄糖苷(kaempferol-3-O-gluciside)、Carthamin、芹菜素(apigenin)、黄芩素(scutellarein)、木犀草素(luteolin)、苯丙氨酸(D-phenylalanine) 12个主要成分的含量[12,25]进行皮尔森相关性分析。
-
基于红花花冠EST转录组文库,结合第三代测序技术[26-29]红花花冠全长转录组数据库筛选得到目的基因序列。在其5'端、3'端分别设计特异性引物。按照2× Phanta Flash Master Mix(Dye Plus)高保真酶(南京诺唯赞公司,中国)说明书进行PCR扩增,扩增片段经EasyPure Quick Gel Extraction Kit胶回收试剂盒(北京全式金公司,中国)说明书操作回收后,连接于pEASY-Blunt Zero Cloning Kit(北京全式金公司,中国)载体上,转化至大肠杆菌T1感受态细胞(北京全式金公司,中国)后,涂布在LBA平板上,恒温培养37 ℃过夜,挑取阳性单克隆菌落[30-31],送至上海生工生物有限公司进行菌液测序。
用ExPASyProtParam工具(http://web.expasy.org/compute/)对目的基因的理论等电点(pI),蛋白分子量(MW)和蛋白分子式进行预测。通过Simple Molecule Architecture Research Tool工具(http://smart.embl-heidelberg.de/)对目的基因编码的蛋白质结构功能域进行分析。使用ProtScale(http://us.Expasy.org/cgi-bin/protscale.pl)以及TMHMM(http://www.cbs.dtu.dk/services/TMHMM/)对蛋白质的亲/疏水性和跨膜区域做出预测。使用SignaIP 4.0(http://www.cbs.dtu.dk/services/SignalP/)预测目的蛋白是否含有信号肽。使用NCBI BLAST(https://blast.ncbi.nlm.nih.gov/Blast.cgi)对筛选出的SDRs基因进行BLAST序列比对。通过Neighbor-Joining相邻节点法构建系统发育进化树,自展分析法进行1000次重复[32-34]。使用PBILYON-GRLAND数据库预测构建蛋白质二级结构模型。蛋白三级结构由Protein Homology/analogy Recognition Engine预测。用WOLFPSORT软件(https://wolfpsort.hgc.jp/)进行亚细胞定位预测。
-
取盛花期新鲜红花根、茎、叶、花冠4个部位的新鲜组织和花期Ⅰ(开花前3 d)、花期Ⅱ(开花当天)、花期Ⅲ(开花后1 d)、花期Ⅳ(开花后3 d)4个花期的新鲜花冠,提取总RNA,合成cDNA第一链后,在靠近5'端处对各个基因设计引物,依据Transtart Top Green qPCR super Mix(北京全式金公司,中国)试剂盒推荐体系,以Ct60s(KJ634810)作为内参标记基因,进行qRT-PCR实验,结果使用2−ΔΔCt的方法进行计算分析[35]。
-
根据CtSDR3的开放阅读框和植物真核表达载体pMT-39序列信息,设计无缝克隆引物。以红花cDNA做模板,使用高保真酶进行PCR反应。产物经胶回收后依无缝克隆试剂盒说明书与经NcoI酶切线性化的pMT-39载体进行重组连接。重组载体转化大肠杆菌T1感受态细胞,挑取阳性克隆菌株扩大培养后抽提质粒,提取的pMT39-CtSDR3质粒用冷冻法转至农杆菌GV3101中。LBK+Rif平板筛选阳性克隆后,取1ml OD600 = 0.8的菌液经6 000 r/min,离心3 min后用1 ml 5%蔗糖溶液重悬,加入Silwet-L 1μl,用注射器注射于红花花柱,套袋避光[35]。
在pMT-39的35 s启动子区域设计5'端特异性引物,在目的基因CtSDR3中设计3'端引物。取T2代新鲜叶cDNA第一链作为模板,2× Easy Taq PCR Mix(北京全式金,中国)推荐体系进行PCR反应,确定是否存在目的条带。采集CtSDR3阳性植株花冠以及pMT-39空载体对照植株的花冠,按照上述的qRT-PCR反应体系评价CtSDR3基因的过表达水平,使用UPLC-Q-TOF/MS 检测CtSDR3过表达组和空载体对照组的黄酮代谢物含量,选择以HSYA为代表性成分的8个黄酮类化合物作为检测对象。
-
根据CtSDR3的开放阅读框及蛋白表达载体pGEX-6p-1以及pET-28a序列信息,设计同源重组克隆引物[34]。以红花花冠cDNA为模板,使用高保真酶进行PCR反应。PCR产物经胶回收后依无缝克隆试剂盒说明书与经XhoI、BamHI酶切线性化的载体pGEX-6p-1以及pET-28a进行重组连接。重组载体转化大肠杆菌T1感受态细胞,挑取阳性菌株克隆扩大培养后抽提质粒,提取的重组质粒用热激法转至大肠杆菌Rosseta(DE3)(上海唯地生物,中国)中。
在20 ml LBA液体培养基中培养至OD600为0.6左右,分2份10 ml菌液各加入终浓度为0.3 mmol/L的IPTG和生理盐水。恒温培养箱中16 ℃,100 r/min继续培养16 h[35-36]。菌液离心弃上清液,用1×PBS缓冲液洗涤两次后重悬。超声破碎仪中40 kW,工作时间5 s,循环间隔时间25 s,共15个循环进行破碎[16],裂解完成后取上清与沉淀15 μl,上样检测。
-
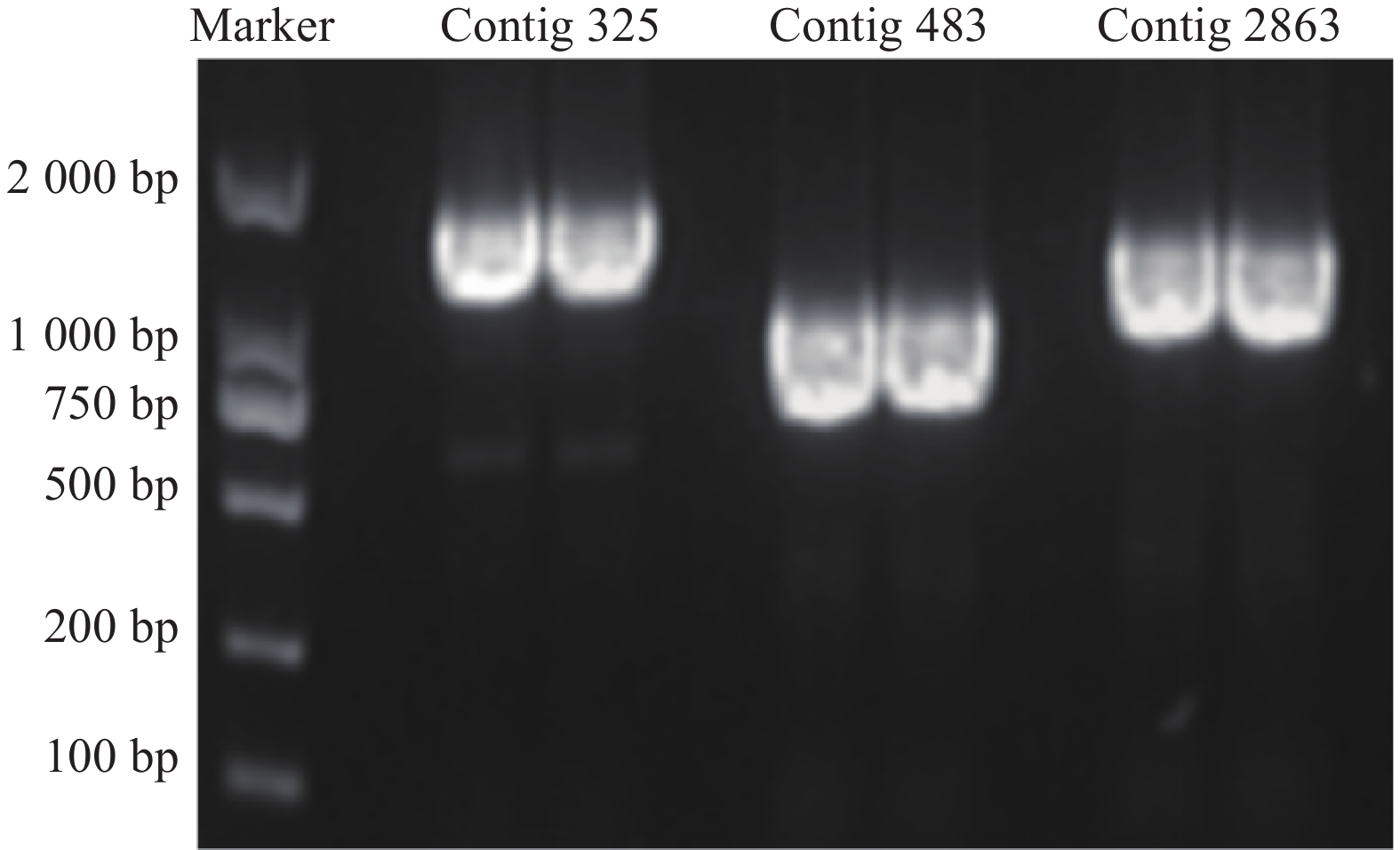
通过分析,得到contig325、contig483、contig2863共3个与HSYA具有强相关性的基因(r>0.85),见图1。
-
3个目的基因序列信息经测序验证结果如下:contig325全长共1523 bp,开放阅读框1341bp,编码446个氨基酸;contig483全长1393 bp,开放阅读框792 bp,编码263个氨基酸;contig2863全长序列1527 bp,开放阅读框1023 bp,编码340个氨基酸。PCR产物电泳结果如图2所示。
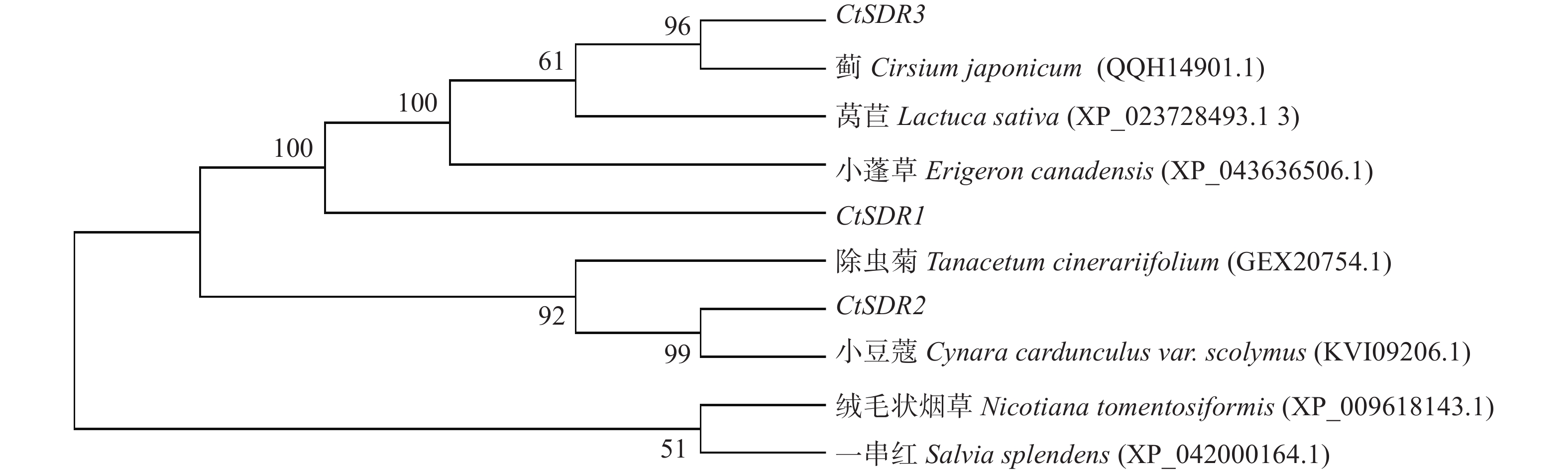
contig325基因编码446个氨基酸,命名为CtSDR1(GenBank登录号:MW792035);Contig483基因编码263个氨基酸,命名为CtSDR2(GenBank登录号:MW792036);Contig2863基因编码339个氨基酸,命名为CtSDR3(GenBank登录号:MW792037)。系统进化树表明CtSDR1与蓟Cirsium japonicum (QQH14901.1)同源性最高;CtSDR2与小蓬草Erigeron canadensis (XP_043636506.1)同源性最高;CtSDR3与小豆蔻Cynara cardunculus var. scolymus (KVI09206.1)同源性最高。Prot-param分析CtSDR1基因所编码的蛋白质分子式C2230H3346N606O639S7,相对分子量为49.2×103,理论等电点pI=9.61;CtSDR2基因所编码的蛋白质分子式C1289H2072N360O379S13,相对分子量为29×103,理论等电点pI=8.63;CtSDR1基因所编码的蛋白质分子式C1691H2614N442O481S9,相对分子量为37.1×103,理论等电点pI=6.80。Prot Scale分析预测CtSDR1、CtSDR2和CtSDR3蛋白为亲水性蛋白,无信号肽属非分泌蛋白。蛋白跨膜性分析显示CtSDR1、CtSDR2和CtSDR3不含有跨膜区域,为非跨膜蛋白。对CtSDR1、CtSDR2和CtSDR3蛋白二级结构的预测显示均属于不规则结构。对CtSDR1、CtSDR2、CtSDR3蛋白质三维结构预测如图3所示。系统进化树如图4所示。亚细胞定位预测显示,CtSDR1、CtSDR2、CtSDR3均可能定位于细胞质。
-
取红花花期的Ⅳ期的红花各个部位进行分析,发现红花花冠内的CtSDR1、CtSDR2、CtSDR3基因表达量从高到低依次均为花冠>叶>茎>根。其中CtSDR1在花冠中的相对表达量约为根中的3倍、而CtSDR2、CtSDR3在花冠中的相对表达量约为根中的4倍。将4个花期的红花花冠进行qRT-PCR分析表明,CtSDR1、CtSDR2、CtSDR3花冠中表达量均随着花冠发育逐渐升高,特别是CtSDR1、CtSDR2、CtSDR3的Ⅳ期花冠对比Ⅲ期花冠的表达量分别提高了7.2倍、2.7倍、2.3倍(图5)。
-
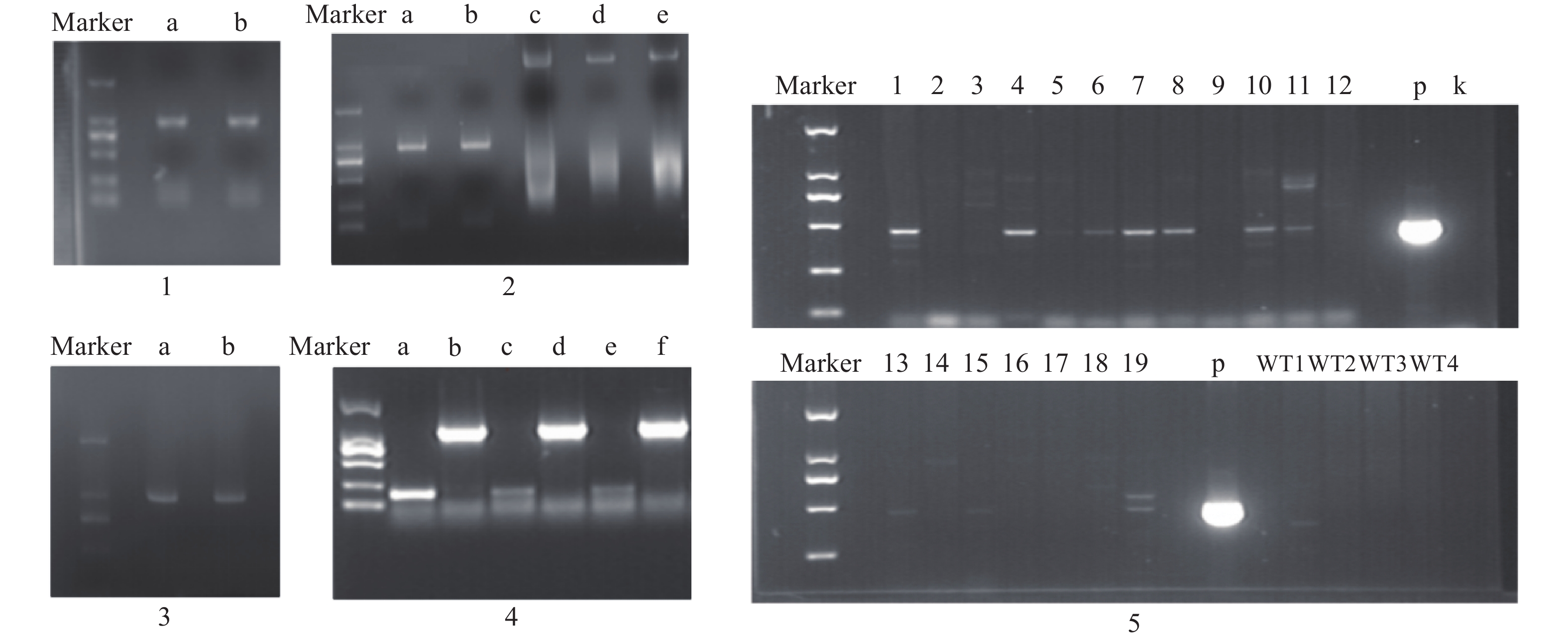
构建真和表达载体并通过PCR鉴定后,我们从19株农杆菌浸染的子代植株中得到5株pMT39-CtSDR3阳性红花植株(图6)。通过qPCR对其CtSDR3基因转录水平进行测定,结果发现阳性红花植株中CtSDR3基因的转录水平得到显著增加,约为空白组株系的2~3倍,CtSDR3的在花冠部位的高表达也证明了研究成功获取CtSDR3过表达红花植株(图7)。通过UPLC-QTOF/MS技术测定阳性转基因红花株系组和空白对照组的目标化合物含量,包括7个红花花冠主要黄酮类化合物及苯丙烷类代谢途径上游关键物质苯丙氨酸(图8),分别为:野黄芩素(scutellarein)、Carthamin、HSYA、山柰酚(kaempferol)、山柰酚-3-O-β-D-葡萄糖苷(kaempferol-3-O-β-D-glucoside)、山柰酚-3-O-β-D-芸香糖苷(kaempferol-3-O-β-rutinoside)、芦丁(rutin)和苯丙氨酸(D-Phenylalanine)。由图8可知,与空白组相比,CtSDR3过表达株系相较于空白组野黄芩素提高了3.6%~9.8%,HSYA提高了7.1%~16.6%,以及苯丙氨酸含量提高了5.5%~15.7%,具有显著性升高。其他化合物含量则有无显著性变化趋势。通过对过表达株系与空白组的含量分析,我们认为CtSDR3基因过表达会引起红花中黄酮类物质的变化,尤其是HSYA含量升高显著。同时,苯丙氨酸代谢途径属于植物重要的次生代谢途径,过表达组引起苯丙氨酸含量的显著上升,上述指标性成分的变化也进一步说明CtSDR3对红花黄酮类化合物次生代谢途径具有一定的影响,但目前我们尚难以判断CtSDR3红花中影响次生代谢产物积累的明确途径。
-
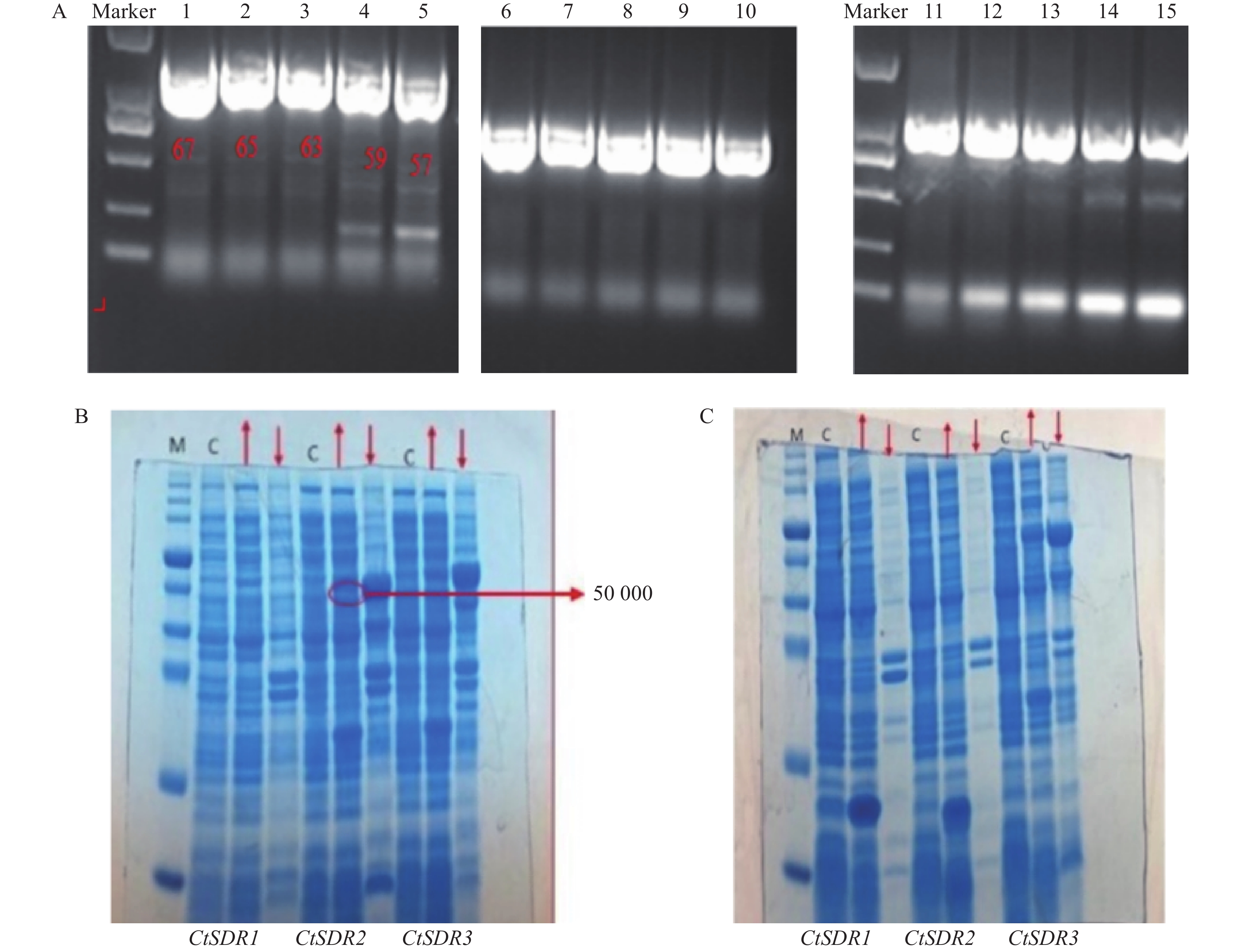
目的片段成功扩增,将目的条带进行胶回收、纯化。CtSDR1、CtSDR1、CtSDR1构建的pGEX-6p-1、pET-28a原核表达载体均有在大肠杆菌内表达,但是CtSDR1-pGEX-6p-1、 CtSDR2-pGEX-6p-1、CtSDR3-pGEX-6p-1、CtSDR1-pET-28、CtSDR2-pET-28a、CtSDR3-pET-28a表达的目的蛋白均形成包涵体,存在于沉淀中。无法进行下一步大量纯化实验,唯有CtSDR2-pGEX-6p-1诱导表达了存在于上清液的目的蛋白,明显可以在上清液中观察到分子量约为50 000的蛋白条带(图9)。
-
越来越多的红花药理学相关研究表明,红花的主要药效物质包括查尔酮类、黄酮醇类等多种黄酮类化合物,其中,查尔酮类HSYA对脑缺血具有保护作用,并且还能抗脑血栓形成以及抗氧化等。研究HSYA的生物合成分途径,对于HSYA的工业化生产具有重要意义。
本研究借助生物学分子技术、结合代谢组分析测定,筛选出3个参与HSYA生物合成途径的关键短链脱氢还原酶基因CtSDR1、CtSDR2和CtSDR3,这3个基因序列具有高度保守性,在不同器官的表达模式均呈现出花冠>叶>茎>根的特点,而且在花冠中的表达量随花冠发育逐渐升高,表明其很有可能参与红花中HSYA等主要药用成分的积累。进一步研究发现,转CtSDR3过表达T2代阳性植株花冠中CtSDR3基因的转录水平增加了2~3倍,次生代谢物HSYA的含量提高了7.1%~16.6%(P<0.05),验证了我们对CtSDR3在红花体内参与黄酮类化合物生物合成功能的推测。本研究中,体外表达CtSDR3蛋白,得到目的蛋白条带,但由于包涵体等原因,蛋白表达和纯化条件仍需要进一步摸索。下一步,我们将对可能起黄酮类生物合成途径的关键SDRs进行深入的生物学特性特别是酶结合位点的研究,为更好地阐释SDRs的生物学功能、利用分子生物育种技术培育高HSYA含量的红花新品种奠定基础。
Characterization and function of short-chain dehydrogenases/reductases in hydroxysafflower yellow A biosynthesis pathway
-
摘要:
目的 探究参与红花黄酮类生物合成途径特别是羟基红花黄色素A(HSYA)关键短链脱氢还原酶基因(SDRs)的功能。 方法 基于红花转录组数据库以及代谢组数据库,筛选参与HSYA生物合成途径的SDRs,用qRT-PCR法分析表达模式。采用无缝克隆技术构建过表达载体,以农杆菌GV3101介导遗传转化云南巍山红花品系,对转基因T2代植株进行阳性验证,并对花冠SDRs的基因表达量进行分析,UPLC-Q-TOF/MS法测定次生代谢物的含量。 结果 筛选出3个参与HSYA生物合成途径的关键短链脱氢还原酶基因CtSDR1、CtSDR2、CtSDR3,表达量从高到低依次为花冠>叶>茎>根。花冠中表达量随花冠发育逐渐升高。对转基因T2代植株进行阳性验证后的花冠进行qRT-PCR分析发现:与空白对照组相比,转CtSDR3过表达T2代阳性植株花冠中CtSDR3基因的转录水平增加了2~3倍,次生代谢物HSYA的含量提高了7.1%~16.6%(P<0.05)。 结论 CtSDR3可能参与了红花中黄酮类化合物特别是HSYA的生物合成,为阐释CtSDR3在HSYA生物合成途径中的功能提供了数据支撑。 Abstract:Objective To explore the function of short-chain dehydrogenases/reductases (SDRs) in safflower flavonoid, especially hydroxysafflower yellow A (HSYA) biosynthesis. Methods SDRs involved in HSYA biosynthesis pathway were screened based on safflower transcriptome database and metabolome database. The expression pattern was analyzed by qRT-PCR. The overexpression vector was constructed by seamless cloning technology, then genetically transformed to the Yunnan Weishan safflower strain by Agrobacterium gv3101. The transgenic T2 generation plants were positively verified, and the gene expression of corolla SDRs was analyzed. The content of secondary metabolites was assayed by UPLC-Q-TOF/MS. Results Three SDRs genes named CtSDR1, CtSDR2 and CtSDR3 involved in HSYA biosynthesis pathway were screened. Their expression in safflower from high to low was corolla > leaf > stem > root. The expression level in corolla increased gradually with corolla development. qRT-PCR analysis of corolla with positive verification of genome insertion sequence showed that the transcription level of CtSDR3 in corolla of T2 positive plants increased by 2~3 times compared with the blank control group, and the content of secondary metabolite HSYA increased by 7.1%~16.6% (P< 0.05). Conclusion CtSDR3 may be involved in the biosynthesis of flavonoids, especially HSYA, in safflower. It provides the support data for explaining the function of CtSDR3 in HSYA biosynthesis pathway. -
Key words:
- short-chain dehydrogenase reductase /
- hydroxysafflor yellow A /
- safflower
-
药品不良反应(ADR)是指正常剂量的药物用于预防、诊断、治疗疾病或调节生理功能时出现的有害的和与用药目的无关的反应[1]。ADR是药品的固有属性,通常所有药品都会或多或少地存在或轻或重的不良反应。据国家不良反应监测年度报告,随着药物种类的增多和用药范围的扩大以及大众对ADR的重视,ADR的报告率逐年上升[2]。 当今人口老龄化的加剧,服用多种药品的人群比例越来越高。有数据表明,药物使用品种数与ADR发生率成正相关。在每日服用超过5种药品的患者中,ADR发生率达25%,这给患者的健康和生命安全带来了严重威胁[3]。为了深入分析后疫情时期ADR发生的特点,本研究对上海交通大学附属第六人民医院南院(我院)2021−2023年的ADR报告进行系统梳理,着重了解ADR的发生情况、影响因素和应对措施,为保障患者安全和合理用药提供科学依据。
1. 资料与方法
1.1 一般资料
回顾性收集我院2021−2023年上报至国家药品不良反应监测中心的ADR报告,共979例。
1.2 调查方法
本研究采用了数据分析和统计描述的方法,数据提取包括患者基本信息(如年龄、性别等)、用药情况(如药物品种、剂量、使用时间等)、不良反应类型及表现(如皮疹、恶心、呕吐等)、治疗及转归情况等。运用Excel软件,对ADR报告进行数据分析。对发生ADR的药物、累积的器官/系统、严重情况等进行分类和统计。对其中缺失数据、不规范数据予以剔除,共收集到符合要求的979例ADR报告。通过计算各类指标的平均值、标准差、频数和百分比等,以反映ADR报告的整体特征和趋势。
2. 结果
2.1 ADR患者基本信息
在979例发生ADR的患者中,男性622例(63.5%),女性357例(36.4%)。发生ADR患者平均年龄为56.8岁,最小年龄为3岁,最大年龄为88岁(见表1)。
表 1 2021−2023年我院ADR患者的年龄及性别分布年龄(岁) 例数(n) 总例数及构成比
[n(%)]男性 女性 ≤10 12 4 16(1.6) 11~20 11 4 15(1.5) 21~30 88 71 159(16.2) 31~40 122 79 201(20.5) 41~50 136 73 209(21.3) 51~60 101 61 162(16.5) 61~70 107 47 154(15.7) 71~80 41 15 56(5.7) 81~90 4 3 7(0.7) 合计 622 357 979(100) 在2021年的354例ADR报告中,男性213例,占比60.17%,女性141例占比39.83%,男女比例为1.51∶1。患者年龄最小者3岁,年龄最大者88岁。各年龄段ADR均有发生,31~40岁和41~50岁是ADR的高发人群,分别占总数的19.21%和23.45%。在2022年302例ADR报告中,男性207例(68.54%),女性95例(31.45%),男女比例为2.19∶1。患者年龄最小者4岁,年龄最大者86岁。各年龄段ADR均有发生,31~40岁和41~50岁是ADR的高发人群,分别占总数的19.86%和25.16%。在2023年323例ADR报告中,男性202例(63.53%),女性121例(37.47%),男女比例为1.71∶1。患者年龄最小者8岁,年龄最大者81岁。各年龄段ADR均有发生, 31~40岁和61~70岁是ADR的高发生人群,分别占总数的23.57%和21.67%。总体而言,2021−2023年我院发生ADR以男性患者居多,且年龄在31~50岁人群发生ADR的概率最高。
2.2 ADR发生严重情况及转归
979例发生ADR中,977例在停用发生ADR的药物后病情有所好转或痊愈,占比为99.8%。2021年报告的354例ADR中353例患者在接受适当的治疗后,病情有所好转或痊愈,占比超过99.7%。仅有1例患者的不良反应未能好转,占0.3%。2022年报告了302例ADR。其中,301例患者在接受适当的治疗后,病情有所好转。2023年报告了323例ADR,所有发生ADR患者均已好转,3年来,该院发生新的ADR合计219例,占比22.4%(见表2)。以上数据表明,2021−2023年我院ADR发生率总体平稳,ADR的转归情况均表现良好。
表 2 2021−2023年我院药品不良反应发生与转归情况年份
(年)总例数
(n)不同程度例数(n) 新发例数
[n(%)]好转或痊愈
例数(n)未好转
例数(n)一般 严重 2021 354 353 1 88(24.9) 353 1 2022 302 301 1 55(18.2) 301 1 2023 323 323 0 76(23.5) 323 0 合计 979 977 2 219(22.4) 977 2 2.3 ADR药物分类
通过对我院2021−2023年的ADR数据分析发现,引发不良反应的药物有135个品种,包括抗菌药物、中药注射剂、维生素类、中成药、镇痛药、脱水药等,其中抗菌药物引发ADR 478例,占比48.8%,排在ADR发生比例的首位:2021年报告168例,占比47.46%、2022年报告146例,占比48.34%、2023年报告146例,占比50.77%;其次是中成药注射剂,占比19.2%,维生素类药物ADR发生率同样不容小觑,发生了73例,占比7.5%,中成药和马破伤风免疫球蛋白分别位列第4、第5,占比分别是7.2%和6.3%(见表3)。
表 3 2021−2023年我院发生ADR的药物分类药品种类 品种数(n) 例数(n) 构成比(%) 抗菌药物 42 478 48.8 中药注射剂 18 188 19.2 维生素类 3 73 7.5 中成药 21 70 7.2 马破伤风免疫球蛋白 3 62 6.3 镇痛药 9 34 3.5 脱水药 3 20 2.0 抗凝血药 3 8 0.8 造影剂 3 7 0.7 降血糖药 6 6 0.6 抗病毒药 3 6 0.6 糖皮质激素 3 5 0.5 其他 18 22 2.2 合计 135 979 100 2.4 发生ADR的抗菌药物种类
进一步对3年间478例发生ADR的抗菌药物进行分类整理,发现头孢呋辛、头孢他啶、头孢替安、头孢唑啉、头孢唑肟、头孢噻肟等头孢菌素类引起的不良反应居多,有214例,占比44.8%;其次是头孢美唑、头孢西丁等头霉素类有120例,占比25.1%;喹诺酮类有59例,占比12.3%。大环内酯类、青霉素类、糖肽类等抗生素类ADR占比虽少,但仍不应轻视(见表4)。
表 4 2021−2023年我院发生ADR抗菌药物品种分布类别 药品 例数(n) 构成比
(%)头孢菌素类 头孢呋辛、头孢他啶、头孢替安、
头孢唑啉、头孢唑肟、头孢噻肟214 44.8 头霉素类 头孢美唑、头孢西丁 120 25.1 喹诺酮类 左氧氟沙星 59 12.3 大环内酯类 阿奇霉素 44 9.2 其他 克林霉素、磷霉素 23 4.8 糖肽类 万古霉素 10 2.1 青霉素类 青霉素 8 1.7 合计 478 100.0 2021年168例抗菌药物ADR事件中,头孢菌素类有77例,占比45.8%;头霉素类有44例,占比26.2%;喹诺酮类有23例,占比13.7%;大环内酯类有4例,占比2.4%。2022年146例抗菌药物ADR事件中,头孢菌素类有56例,占比38.4%;头霉素类有38例,占比26%;喹诺酮类有27例,占比18.5%;大环内酯类有19例,占比13%。2023年164例抗菌药物ADR事件中,头孢菌素类有81例,占比49.4%;头霉素类有38例,占比23.2%;喹诺酮类有9例,占比5.5%;大环内酯类有21例,占比12.8%。总体而言,2021−2023年我院抗菌药物ADR中头孢菌素类仍有较高的占比,需持续关注。
2.5 发生ADR的给药途径
通过对3年间979例发生ADR患者的给药途径统计发现,静脉滴注导致的ADR患者763例,占比77.9%;口服导致的ADR患者128例,占比13.1%;外用导致的ADR患者4例,占比0.4%(见表5)。其中,2021年发生的ADR患者中,静脉滴注导致的ADR患者296例,占比83.6%;口服导致的ADR患者27例,占比7.6%;2022年发生的ADR患者中,静脉滴注导致的ADR患者231例,占比76.5%;口服导致的ADR患者40例,占比13.2%;2023年发生的ADR患者中,静脉滴注导致的ADR患者236例,占比73.1%;口服导致的ADR患者61例,占比18.9%。表明2021−2023年静脉滴注是导致ADR的主要给药途径,虽然静脉滴注给药效果显著,但安全性问题容易被忽视,仍需给予关注。
表 5 2021−2023年我院发生ADR的给药途径年份(年) 不同给药途径例数(n) 静脉滴注 口服 外用 其他 2021 296 27 0 31 2022 231 40 3 28 2023 236 61 1 25 合计 763 128 4 84 2.6 累及器官/系统及主要临床表现
通过分析我院在2021−2023年ADR患者的临床表现,发现542例患者在皮肤方面有不良反应,临床表现是皮疹、瘙痒、斑丘疹、荨麻疹、潮红等,占比55.4%;168例患者主要在消化系统方面有不良反应,临床表现为恶心、呕吐、腹泻、腹痛、胃痛、腹胀、胃部不适、食欲减退等,占比为17.2%;168例患者主要在呼吸系统方面有不良反应,临床表现为呼吸困难、胸闷、胸痛、气促、憋气等,占比为10.3%;59例患者有局部不良反应,临床表现为静脉炎、局部红肿、血管性水肿、面部水肿等,占比为6%(见表6)。2021年ADR损害以皮肤及其附件为多见,有216例,占比61.02%;呼吸系统方面有不良反应的患者有61例,占比17.2%。2022年ADR损害同样以皮肤及其附件为多见,有180例,占比59.6%;呼吸系统方面有不良反应的患者有35例,占比11.6%。2023年ADR累及皮肤及其附件占比为45.2%,较前略有下降;呼吸系统方面有不良反应的患者有72例,占比22.3%。以上数据表明2021−2023年我院ADR累及器官/系统及主要临床表现为皮肤及附件损害和呼吸系统方面,但循环系统、全身性损害、神经系统等临床不良反应患者仍需关注。
表 6 2021−2023年我院发生ADR累及器官或系统及其临床表现累及器官/系统 主要临床表现 例数(n) 构成比(%) 皮肤及附件损害 皮疹、瘙痒、斑丘疹、荨麻疹、潮红、血管性水肿、面部水肿等 542 55.4 消化系统 恶心、呕吐、腹泻、腹痛、胃痛、腹胀、胃部不适、食欲
减退等168 17.2 呼吸系统 呼吸困难、胸闷、胸痛、气促、憋气等 101 10.3 局部反应 静脉炎、局部红肿等 59 6.0 神经系统 头晕、头痛、眩晕、抽搐、失眠、焦虑等 53 5.4 全身性损害 寒颤、发热、过敏性休克、水肿、晕厥、疼痛 36 3.7 循环系统 心慌、心悸、心动过速/过缓、紫绀等 20 2.0 总计 979 100.0 3. 讨论
在过去的几年中,ADR问题一直是医疗领域关注的焦点之一。这些不良反应不仅对患者健康产生负面影响,还会增加医疗成本和资源消耗。因此,对药物不良反应进行监测和分析,对于保障患者安全和促进合理用药具有重要意义。
本文报告了我院2021−2023年期间发生的979例ADR,其中2022年ADR上报数量偏少,其中4−6月ADR上报数量明显低于同期,推测可能与2022年当时上海的疫情有关。
3.1 ADR与患者性别、年龄的关系
流行病学调查发现中国高血压和脑卒中患病率均男性高于女性[4-5]。我国及全球癌症统计均显示,除女性特有癌症外,通常男性癌症发生率高于女性[6-8]。男性发病几率大于女性提示男性用药人群多,发生ADR的概率比例高,我们的ADR分析也佐证了这一点。从发生ADR的年龄分布来看,好发生ADR的年龄区间在21~70岁,尤其是30~50岁中青年区间。一般认为幼龄儿童和老年人由于器官功能不全容易发生ADR,但我们的数据却发现中青年人群ADR发生率居高,推测可能与此年龄段人群依从性低,遵医嘱率低有关,这一结果与张成栋等[9]的ADR分析报告相似。
3.2 ADR与药物类别的关系和应对措施
从本次调查发生ADR的药物分类来看,排名较为靠前的是抗菌药物、中药注射剂、维生素类药物、中成药及马破伤风免疫球蛋白。随着近年来新型冠状病毒肺炎(COVID-19)、流行性感冒等疾病的大爆发,病毒合并细菌感染人群急剧增多,抗菌药物使用也在不断增多,临床调查发现,抗菌药物ADR的发生率有所增高[10]。过度使用和滥用抗菌药物会导致耐药菌株的产生和传播,增加治疗难度和医疗成本。随着我国对抗菌药物合理使用重视程度不断提高,临床对抗菌药物的监管亦日益增强。在抗菌药物使用过程中,医生需要严格控制其适应证和用法用量,根据患者的病情和药敏试验结果,合理选择抗菌药物,避免不必要联合用药和长期使用。必要时进行血药浓度监测以加强用药安全,避免ADR的发生[11]。
3.3 ADR发生与给药途径的关系
本次调查的979例ADR中,注射液导致的ADR 763例,占比77.9%;口服导致的ADR 128例,占比13.1%。这可能与我院上报ADR多为住院患者有关,因静脉给药是当前住院患者药物治疗的主要途径。静脉给药无首过作用,药物可直接进入血液循环,生物利用度高,可迅速发挥药效,而受到临床青睐,患者也相信静脉给药比口服给药更有效果,但静脉给药的安全性却容易被忽视[12]。我们的研究再次提醒医务人员和患者,尽量不选择静脉给药,在通过静脉滴注给药时,为减少ADR发生,应严格控制滴注速度,防止因滴速过快而短时间内血药浓度过高导致ADR发生;另外避免大剂量超剂量给药,这在临床较为普遍,尤其是头孢类抗菌药物,由于2次/d的给药频次给门急诊患者带来不便,许多医生将1 d的剂量1次给药,这大大增加了药物峰浓度水平而导致ADR的发生。值得注意的是,分析3年来ADR发生与给药途径差异发现,静脉给药导致的ADR逐年下降,从83.6%下降到73.1%。推测可能与近年来门急诊的输液率下降有关。近年来,各大医院严格控制门急诊患者的输液比率,多数医院已基本取消门诊患者输液,从而也减少了部分输液所致的ADR。
3.4 ADR 累及的系统/组织及临床表现
本次调查的979例ADR中,大部分临床表现比较轻微。皮肤损害、消化系统损害和呼吸系统损害占前3位。但仔细比较3年来ADR累及组织器官的差异发现,累及皮肤系统的占比在逐年下降,而累及呼吸系统的ADR在2023年占比最高,为22.3%,较2022年翻一番。分析可能是2022年底COVID-19疫情全面放开后,新型冠状病毒攻击了全国大多数居民,导致2023年无论是新型冠病状病毒所致感染还是其他基础疾病等用药增加,而新型冠状病毒攻击的最主要部位就是呼吸系统,从而导致相当一部分居民呼吸系统功能受损,进而使得药物相关的呼吸系统ADR居多[13-14]。已有不少研究报道COVID-19治疗中常用药物的呼吸系统不良反应。如某些抗病毒药物和免疫抑制剂可能引发肺炎、肺纤维化等严重呼吸系统反应。此外,亦有研究发现COVID-19疫情期间,哮喘及心血管疾病的复发而致平喘、抗心血管药物使用增加,这些药物对呼吸系统的不良反应比例上升[15-16]。
3.5 ADR的监测展望
面对日益增加的ADR报道,尤其是新药不断上市的今天,新的和严重的ADR发生问题也日益突出。因此ADR的监测和预防成为了医学研究的重要课题。陈君恒等[17]利用消息传递神经网络(MPNN)和TransE模型进行建模,通过交叉压缩单元为共享,建立了可实现对不良药物检测的多任务消息传递神经网络(MT-MPNN)模型。O’Leary等[18]对新药开发过程中的ADR进行了研究,研发以人工智能为基础的ADR监控体系,在海量的临床试验中对ADR进行实时监控与预警,并能够给临床医师带来病人对某一类药品可能发生ADR的危险程度预警,进而帮助医师做出更为个体化的用药策略。除了开发智能系统外,人工干预如临床药师的干预同样对提高ADRs的上报质量和数量至关重要[19]。
ADR的发生是一个复杂的过程,涉及到多个因素和环节,需要我们从多个角度进行研究和监测,以提高药物治疗的安全性和有效性。临床医务工作者应加强对ADR的发生情况、特点和规律的了解,对严重或者特殊的ADR类型提高警惕,及时发现和处理。应特别注意抗菌药物的合理使用,避免中药注射剂等不必要的使用,给药方案个体化,关注患者输液反应,加强用药指导和用药服务,纠正不合理用药的情况,提高用药的合理性和安全性。
-
图 6 真核表达载体构建及阳性鉴定电泳图
注:1. CtSDR3基因开放阅读框(ORF)区扩增产物电泳图,a、b泳道均为CtSDR3基因ORF区克隆PCR产物;2. 真核表达载体pMT-39载体酶切产物电泳图,a、b泳道为CtSDR3 PCR产物,c泳道为pMT-39载体,d、e泳道为pMT-39线性化载体;3. pMT39-CtSDR3重组载体阳性转化子鉴定电泳图,a、b泳道为阳性转化子菌液PCR产物;4. pMT39-CtSDR3质粒转化农杆菌GV3101,a、c和e泳道为空白对照组,b、d和f泳道为阳性克隆菌液PCR产物;5. 红花pMT39-CtSDR3阳性转化植株鉴定PCR产物电泳图,1~19为待鉴定植株,p为pMT39-CtSDR3质粒,k为空白组,WT为野生型红花植株
-
[1] 郭美丽, 张汉明, 张美玉. 红花本草考证[J]. 中药材, 1996(4):202. [2] TU Y H, XUE Y R, GUO D D, et al. Carthami Flos: a review of its ethnopharmacology, pharmacology and clinical applications[J]. REV BRAS FARMACOGN,2015,25(5):553-566. doi: 10.1016/j.bjp.2015.06.001 [3] WANG Y Q, ZHANG S S, NI H L, et al. Autophagy is involved in the neuroprotective effect of nicotiflorin[J]. J Ethnopharmacol,2021,278:114279. doi: 10.1016/j.jep.2021.114279 [4] 贾鑫磊, 何贝轩, 郭丹丹, 等. 红花花冠伸长相关基因CtXTH1的特征与功能研究[J]. 药学学报, 2019, 54(6):1132-1140. [5] 张戈. 红花的化学成分及品质评价[D]. 上海: 第二军医大学, 2001. [6] 张戈, 郭美丽, 张汉明, 等. 不同种质红花药材的高效液相色谱法指纹图谱研究[J]. 第二军医大学学报, 2006, 27(3):280-283. doi: 10.3321/j.issn:0258-879X.2006.03.012 [7] 丁丽丽, 段陈平, 李芳, 等. 红花不同采收期及不同部位中羟基红花黄色素A及山奈素的含量变化[J]. 沈阳药科大学学报, 2015, 32(1):65-69. [8] 杨捷威, 吴婷婷, 郭美丽. 红花组织培养的研究进展[J]. 药学服务与研究, 2012, 12(1):58-62. [9] HUANG L L, YANG X, SUN P, et al. The first Illumina-based de novo transcriptome sequencing and analysis of safflower flowers[J]. PLoS One,2012,7(6):e38653. doi: 10.1371/journal.pone.0038653 [10] COSGROVE D J. New genes and new biological roles for expansins[J]. Curr Opin Plant Biol,2000,3(1):73-78. doi: 10.1016/S1369-5266(99)00039-4 [11] CHENG H, LI L L, XU F, et al. Expression patterns of an isoflavone reductase-like gene and its possible roles in secondary metabolism in Ginkgo biloba[J]. Plant Cell Rep,2013,32(5):637-650. doi: 10.1007/s00299-013-1397-2 [12] MIN T, KASAHARA H, BEDGAR D L, et al. Crystal structures of pinoresinol-lariciresinol and phenylcoumaran benzylic ether reductases and their relationship to isoflavone reductases[J]. J Biol Chem,2003,278(50):50714-50723. doi: 10.1074/jbc.M308493200 [13] 崔扬, 冯彦辉, 陈众峰, 等. 玉米短链脱氢酶基因IDP2557的克隆及其抗旱功能鉴定[J]. 分子植物育种, 2019, 17(22):7300-7305. [14] 王化冰. 黄瓜短链脱氢酶SDR110C家族的生物信息学分析及CsSDR110C14的遗传转化[D]. 泰安: 山东农业大学, 2014. [15] BRAY J E, MARSDEN B D, OPPERMANN U. The human short-chain dehydrogenase/reductase (SDR) superfamily: a bioinformatics summary[J]. Chem Biol Interact,2009,178(1-3):99-109. doi: 10.1016/j.cbi.2008.10.058 [16] JÖRNVALL H, HEDLUND J, BERGMAN T, et al. Superfamilies SDR and MDR: from early ancestry to present forms. Emergence of three lines, a Zn-metalloenzyme, and distinct variabilities[J]. Biochem Bioph Res Commun,2010,396(1):125-130. doi: 10.1016/j.bbrc.2010.03.094 [17] KALLBERG Y, OPPERMANN U, PERSSON B. Classification of the short-chain dehydrogenase/reductase superfamily using hidden Markov models[J]. FEBS Journal,2010,277(10):2375-2386. doi: 10.1111/j.1742-4658.2010.07656.x [18] OPPERMANN U, FILLING C, HULT M, et al. Short-chain dehydrogenases/reductases (SDR): the 2002 update[J]. Chemico-Biological Interactions,2003,143-144:247-253. doi: 10.1016/S0009-2797(02)00164-3 [19] WU Y, WANG Y, MI X F, et al. The QTL GNP1 encodes GA20ox1, which increases grain number and yield by increasing cytokinin activity in rice panicle meristems[J]. PLoS Genet,2016,12(10):e1006386. doi: 10.1371/journal.pgen.1006386 [20] GUO T, CHEN K, DONG N Q, et al. GRAIN SIZE AND NUMBER1 negatively regulates the OsMKKK10-OsMKK4-OsMPK6 cascade to coordinate the trade-off between grain number per panicle and grain size in rice[J]. Plant Cell,2018,30(4):871-888. doi: 10.1105/tpc.17.00959 [21] 靳甜甜. 植物次生代谢途径中黄酮类化合物的生物合成研究[D]. 保定: 河北农业大学, 2014. [22] LEE J, BURNS T H, LIGHT G, et al. Xyloglucan endotransglycosylase/hydrolase genes in cotton and their role in fiber elongation[J]. Planta,2010,232(5):1191-1205. doi: 10.1007/s00425-010-1246-2 [23] SHAO M Y, WANG X D, NI M, et al. Regulation of cotton fiber elongation by xyloglucan endotransglycosylase/hydrolase genes[J]. Genet Mol Res,2011,10(4):3771-3782. doi: 10.4238/2011.October.27.1 [24] 谭政委, 鲁丹丹, 李磊, 等. 红花二氢黄酮醇4-还原酶基因CtDFR1的克隆与分析[J]. 分子植物育种, 2021:1-12. [25] XU F, DENG G, CHENG S, et al. Molecular cloning, characterization and expression of the phenylalanine ammonia-lyase gene from Juglans regia[J]. Molecules,2012,17(7):7810-7823. doi: 10.3390/molecules17077810 [26] SHOJI T, WINZ R, IWASE T, et al. Expression patterns of two tobacco isoflavone reductase-like genes and their possible roles in secondary metabolism in tobacco[J]. Plant Mol Biol,2002,50(3):427-440. doi: 10.1023/A:1019867732278 [27] 朱惠, 杨丽涛, 李杨瑞. 基于三代测序技术的转录组学研究[J]. 生物技术报, 2014(11):130-137. [28] PETIT P, GRANIER T, D'ESTAINTOT B L, et al. Crystal structure of grape dihydroflavonol 4-reductase, a key enzyme in flavonoid biosynthesis[J]. J Mol Biol,2007,368(5):1345-1357. doi: 10.1016/j.jmb.2007.02.088 [29] O'REILLY C, SHEPHERD N S, PEREIRA A, et al. Molecular cloning of the a1 locus of Zea mays using the transposable elements En and Mu1[J]. EMBO J,1985,4(4):877-882. doi: 10.1002/j.1460-2075.1985.tb03713.x [30] 郝爱平. 植物二氢黄酮醇4-还原酶的生物信息学分析[J]. 江苏农业科学, 2014, 42(6):30-34. doi: 10.3969/j.issn.1002-1302.2014.06.009 [31] ISHIDA T, HATTORI S, SANO R, et al. Arabidopsis transparent testa glabra2 is directly regulated by r2r3 myb transcription factors and is involved in regulation of glabra2 transcription in epidermal differentiation[J]. Plant Cell,2007,19(8):2531-2543. doi: 10.1105/tpc.107.052274 [32] ZHANG F, GONZALEZ A, ZHAO M Z, et al. A network of redundant bHLH proteins functions in all TTG1-dependent pathways of Arabidopsis[J]. Development,2003,130(20):4859-4869. doi: 10.1242/dev.00681 [33] 郭丹丹. 红花黄酮类成分生物合成途径关键基因的特征和功能研究[D]. 上海: 海军军医大学, 2019. [34] GRÄFF M, BUCHHOLZ P C F, STOCKINGER P, et al. The Short-chain Dehydrogenase/Reductase Engineering Database (SDRED): a classification and analysis system for a highly diverse enzyme family[J]. Proteins,2019,87(6):443-451. doi: 10.1002/prot.25666 [35] STAVRINIDES A K, TATSIS E C, DANG T T, et al. Discovery of a short-chain dehydrogenase from Catharanthus roseus that produces a new monoterpene indole alkaloid[J]. Chembiochem,2018,19(9):940-948. doi: 10.1002/cbic.201700621 [36] ZHOU Y, ZHANG L, GUI J D, et al. Molecular cloning and characterization of a short-chain dehydrogenase showing activity with volatile compounds isolated from Camellia sinensis[J]. Plant Molecular Biology Reporter,2015,33(2):253-263. doi: 10.1007/s11105-014-0751-z -




 下载:
下载:
 下载:
下载: